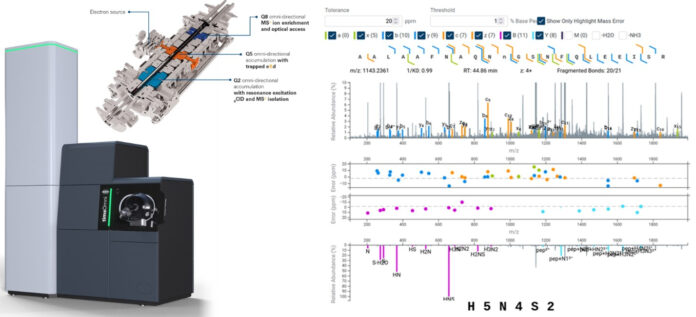
ST. LOUIS — Bruker has announced new advancements to its timsOmni mass spectrometry platform, expanding what the company describes as Functional Proteomics 2.0 and enabling deeper analysis of disease biology, drug targets, and therapeutic mechanisms.
The updates extend timsOmni workflows beyond traditional protein identification toward detailed characterization of biologically active proteoforms and post-translational modification–resolved peptide variants. New software releases across Bruker’s ProteoScape, OmniScape, and GlycoScape platforms support database-independent PTM discovery, high-confidence proteoform identification, and electron-based dissociation–enabled glycoproteomics.
The company said the enhanced capabilities allow researchers to investigate complex molecular pathways and mechanisms of action that are often difficult to resolve with conventional proteomics approaches. In drug discovery and development, the workflows are designed to support analysis of small-molecule targets as well as biologics such as therapeutic antibodies, antibody-drug conjugates, and multispecific therapies, including their modification patterns and glyco-heterogeneity.
The latest OmniScape 2026b release strengthens top-down proteoform analysis by integrating a new algorithm that enables large-scale sequencing and confident annotation of critical post-translational modifications linked to disease signaling and treatment response. According to Bruker, this allows researchers to shift from protein group summaries to proteoform-centric insights that more accurately reflect biological and pathological states.
In biotherapeutics research, the software also supports verification of expected protein sequences and detection of non-canonical modifications or sequence variants that may influence safety or efficacy.
Bruker is also expanding glycoproteomics capabilities by adding support for trapped electron-based dissociation data within GlycoScape. The approach preserves fragile glycan structures while enabling precise localization and topology analysis. Interactive visualization tools allow researchers to validate glycopeptide results directly within annotated mass spectra.
“The timsOmni gives scientists access to electron-based fragmentation that can resolve complex post-translational modifications and proteoforms with outstanding fidelity,” said Professor Yehia Mechref, director of the Texas Tech University Center for Biotechnology & Genomics. “The new software capabilities support deeper understanding of disease biology and open new avenues for target discovery and translational research.”
In addition, Bruker is introducing an AI-enhanced de novo peptide sequencing workflow in ProteoScape 2026b. The new approach combines an artificial intelligence scoring model trained on more than seven million mass spectra with established dynamic programming methods to enable accurate, database-independent peptide sequencing from high-resolution data.
The company said this capability addresses growing demand for proteomics approaches that do not rely solely on reference databases, particularly in fields such as metaproteomics and immunopeptidomics, including neoantigen discovery in immuno-oncology.
“Researchers are increasingly studying biological systems where reference databases alone do not capture the true molecular diversity,” said Bin Ma, founder of Rapid Novor. “Our collaboration has enabled AI-driven sequencing approaches that deliver the speed, confidence, and precision needed for next-generation de novo peptide discovery.”
Bruker said the combined advances position timsOmni as a key platform for illuminating previously inaccessible aspects of molecular biology and accelerating research across disease, diagnostics, and drug development.